A novel machine-learning algorithm used with MRI can harmonize brain volumetric data of patients undergoing Alzheimer's disease assessment gathered from different scanners, researchers have reported.
"When evaluated on brain MRI marker data from participants along the Alzheimer's disease spectrum, our new model outperformed the other approaches we tested on both seen and unseen [i.e., scanners not included in the initial training of the model] scanners," wrote a team led by Damian Archetti, PhD, of IRCCS Istituto Centro San Giovanni di Dio Fatebenefratelli in Brescia, Italy. The findings were published December 18 in Radiology: Artificial Intelligence.
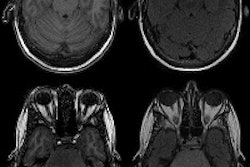
Structural MR imaging, such as T1-weighted exams, are commonly used in memory clinics for diagnosing Alzheimer's disease and differentiating Alzheimer's from other types of dementias, the group explained. Radiologists visually interpret these exams, but these interpretations are subjective and "prone to intrarater and interrater variability," it noted. Using quantitative imaging markers culled from brain volumetric data shows promise for improving readers' diagnostic confidence, and there are AI algorithms that track these. But differences in MRI acquisition protocols and scanners can affect the consistency of brain volumetry assessment.
Neuroharmony is a relatively new tool for harmonizing volumetric data from images taken using new or unseen MRI scanners. It uses image quality metrics (IQM) as predictors to remove scanner-related effects in brain-volumetric data using random forest regression and has been tested on data from cognitively healthy people; however, it has not been tested on data from patients with neurodegenerative diseases, the researchers explained.
To this end, Archetti and colleagues developed an extended version of Neuroharmony to track interactions between Alzheimer's disease pathology and image quality metrics in 20,864 individuals with and without cognitive impairment. These data spanned 11 prospective and retrospective study cohorts and 43 scanners. The group then evaluated the expanded algorithm's ability to remove scanner-related variations in brain volumes (that is, to track marker concordance between scanners) while continuing to delineate disease-related signals.
The investigators found that marker concordances between scanners were significantly better (p < 0.001) compared with data previously processed without the algorithm. They also found that the expanded model showed higher concordance than the original model (0.75 compared with 0.7, with 1 as reference) and better-preserved disease signals (area under the receiver operating curve, or AUC, -0.006 compared with -0.091). Finally, the group reported that marker concordance was better in scanners used to train the algorithm (> 0.97) than in those not used for this purpose (< 0.79) -- a result that emerged independently of study participants' cognitive status.
The study results show promise, but further research is needed, according to the authors.
"Further validation using different processing pipelines and evaluation criteria would be essential for clinical use of the model in applications related to cognitive decline, such as memory clinics and clinical trials of new interventions for neurodegenerative diseases," they concluded.
The complete study can be found here.



![Overview of the study design. (A) The fully automated deep learning framework was developed to estimate body composition (BC) (defined as subcutaneous adipose tissue [SAT] in liters; visceral adipose tissue [VAT] in liters; skeletal muscle [SM] in liters; SM fat fraction [SMFF] as a percentage; and intramuscular adipose tissue [IMAT] in deciliters) from MRI. The fully automated framework comprised one model (model 1) to quantify different BC measures (SAT, VAT, SM, SMFF, and IMAT) as three-dimensional (3D) measures from whole-body MRI scans. The second model (model 2) was trained to identify standardized anatomic landmarks along the craniocaudal body axis (z coordinate field), which allowed for subdividing the whole-body measures into different subregions typically examined on clinical routine MRI scans (chest, abdomen, and pelvis). (B) BC was quantified from whole-body MRI in over 66,000 individuals from two large population-based cohort studies, the UK Biobank (UKB) (36,317 individuals) and the German National Cohort (NAKO) (30,291 individuals). Bar graphs show age distribution by sex and cohort. BMI = body mass index. (C) After the performance assessment of the fully automated framework, the change in BC measures, distributions, and profiles across age decades were investigated. Age-, sex-, and height-adjusted body composition reference curves were calculated and made publicly available in a web-based z-score calculator (https://circ-ml.github.io).](https://img.auntminnie.com/mindful/smg/workspaces/default/uploads/2026/05/body-comp.XgAjTfPj1W.jpg?auto=format%2Ccompress&fit=crop&h=100&q=70&w=100)


![Overview of the study design. (A) The fully automated deep learning framework was developed to estimate body composition (BC) (defined as subcutaneous adipose tissue [SAT] in liters; visceral adipose tissue [VAT] in liters; skeletal muscle [SM] in liters; SM fat fraction [SMFF] as a percentage; and intramuscular adipose tissue [IMAT] in deciliters) from MRI. The fully automated framework comprised one model (model 1) to quantify different BC measures (SAT, VAT, SM, SMFF, and IMAT) as three-dimensional (3D) measures from whole-body MRI scans. The second model (model 2) was trained to identify standardized anatomic landmarks along the craniocaudal body axis (z coordinate field), which allowed for subdividing the whole-body measures into different subregions typically examined on clinical routine MRI scans (chest, abdomen, and pelvis). (B) BC was quantified from whole-body MRI in over 66,000 individuals from two large population-based cohort studies, the UK Biobank (UKB) (36,317 individuals) and the German National Cohort (NAKO) (30,291 individuals). Bar graphs show age distribution by sex and cohort. BMI = body mass index. (C) After the performance assessment of the fully automated framework, the change in BC measures, distributions, and profiles across age decades were investigated. Age-, sex-, and height-adjusted body composition reference curves were calculated and made publicly available in a web-based z-score calculator (https://circ-ml.github.io).](https://img.auntminnie.com/mindful/smg/workspaces/default/uploads/2026/05/body-comp.XgAjTfPj1W.jpg?auto=format%2Ccompress&fit=crop&h=112&q=70&w=112)