MR imaging has revealed the brain's waste-clearing pathways (that is, perivascular spaces, or fluid-filled structures along arteries and veins) for the first time, investigators have reported.
A team led by lead author Erin Yamamoto, MD, and senior author Juan Piantino, MD, both of Oregon Health & Science University (OHSU) in Portland found that "a well-functioning glymphatic system efficiently carries waste proteins toward veins exiting the brain." The results were published October 7 in the Proceedings of the National Academy of Sciences.
"Nobody has shown it before now," Piantino said in a statement released by OHSU. "I was always skeptical about it myself, and there are still a lot of skeptics out there who still don’t believe it. That’s what makes this finding so remarkable."
Researchers have long held a theory that the network of brain pathways clear metabolic proteins that otherwise could build up and lead to dementia and/or Alzheimer's disease. But this network has not been definitively revealed, the team noted.
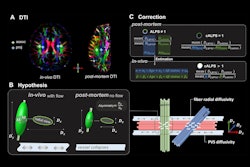
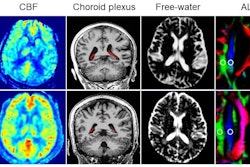
To do so, Piantino and colleagues conducted a study that included five patients who underwent neurosurgery between 2020 and 2023. All of these were injected with a gadolinium-based inert contrasting agent that was used with a fluid-attenuated inversion recovery (FLAIR) MRI exam to track cerebrospinal fluid flowing throughout the brain at 12, 24, and 48 hours after surgery.
These MR images showed the fluid moving along perivascular spaces, "definitively revealing the presence of an efficient waste-clearance system within the human brain," the group wrote.
The findings underscore the importance of lifestyle measures to improve sleep quality, such as maintaining a regular sleep schedule and avoiding screen time before bed in order to support the brain's nighttime waste-clearing.
"People thought these perivascular spaces were important, but it had never been proven," Piantino said in the OHSU statement. "Now it has."
The complete study can be found here.