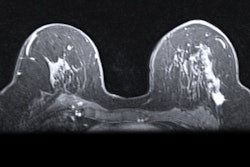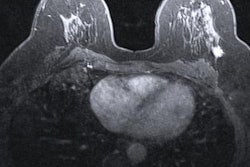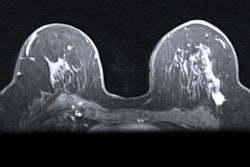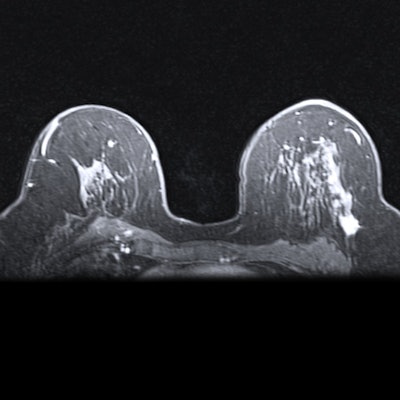
A hybrid artificial intelligence (AI) model that incorporates dynamic contrast-enhanced MRI (DCE-MRI) images acquired prior to chemotherapy accurately predicts complete treatment response in breast cancer patients, researchers have found.
A team led by Priyanka Khanna from the National Institute of Technology Raipur in India found near-perfect accuracy in both hold-out and 10-fold cross-validations with their machine-learning model, which they suggest could help with treatment strategy for those who may not show complete response to conventional strategies. The findings were published December 11 in Measurement.
"Our early response model has the potential to identify the non-responder patients undergoing therapy, thus minimizing toxicity and facilitating alternate treatment plans," Khanna and co-authors wrote.
Artificial intelligence (AI) continues to show potential in breast imaging, whether it be detecting suspicious findings on initial imaging or forecasting treatment response. Early detection of breast cancer leads to improved prognosis via better treatment strategies and can lead to saved time and costs.
Neoadjuvant chemotherapy is typically used for stage II or III breast cancer to reduce tumors and avoid mastectomy, but some patients do not respond well to this method. DCE-MRI is typically preferred to predict therapy response, with proponents highlighting its ability to accurately measure tumor dimensions.
Khanna and colleagues wanted to explore the efficacy of an automated system to assist radiologists in predicting chemotherapy response by using baseline breast MRI tumor datasets in their model. The group integrated DCE-MRI images acquired before the start of chemotherapy into a pre-trained convolutional neural network (CNN) with machine learning, using ResNet-50 and ResNet-18 for feature extraction.
The researchers tested the model on data from 64 women who received chemotherapy for breast cancer treatment.
They found that the hybrid model had an accuracy of 99.8% and yielded an area under the receiving operator curve (AUROC) of 1 for the hold-out validation. For 10-fold cross-validation, accuracy was 99.3% while the AUROC was 0.99.
The study authors wrote that they intend to extend their work with larger and other datasets, as well as evaluate deep learning-based segmentation methods for detecting regions of interest and other areas of optimization.
"For easy access by radiologists and oncologists, the deep-learning model can be integrated into an interactive web application," they noted.