An artificial intelligence (AI) algorithm was able to preoperatively identify lymph node metastasis on MRI in patients with cervical cancer, potentially enabling some patients to avoid unnecessary surgery, according to research published online July 24 in JAMA Network Open.
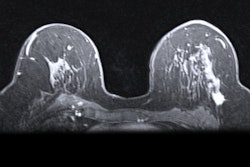
A team of researchers led by Dr. Qingxia Wu, PhD, of the Chinese Academy of Sciences in Beijing developed an AI model that combined a deep-learning algorithm with assessment of lymph-node status on MRI. In testing, the hybrid model yielded an area under the curve (AUC) of 0.933 for detecting lymph node metastases on MRI scans.
"Findings from this study suggest that deep learning can be used as a preoperative noninvasive tool for diagnosing lymph node metastasis in cervical cancer," the authors wrote.
Accurate preoperative identification of lymph node status in cervical cancer may avoid unnecessary surgical intervention and improve treatment planning, according to the researchers. The traditional method of relying on the size of lymph nodes on MRI offers limited sensitivity for diagnosing metastases, however.
Although extracting radiomic features from MRI has shown potential to improve the modality's sensitivity, these features require time-consuming delineation and might not be adaptive to specific clinical issues, according to the authors. As a result, Wu and colleagues sought to utilize deep learning to develop a preoperative, noninvasive tool for diagnosing lymph node metastases on MRI.
"Because the [deep-learning] model can mine high-dimensional information from MRI and clinical features can reflect tumor information from clinicopathologic aspects, we developed a hybrid model to combine information from these sources to explore whether they can be complementary," they wrote.
The researchers developed and validated the models on a primary cohort of 338 cervical cancer patients, including 218 patients from Sun Yat-sen University Cancer Center in Guangzhou, China, and 120 patients from Henan Provincial People's Hospital in Zhengzhou, China. Next, they tested the algorithms on an independent validation dataset of 141 patients from Yunnan Cancer Hospital in Kunming, China.
After testing various combinations of models, the researchers found that the best was a hybrid model that integrated a deep-learning model with MRI-reported lymph node status. A positive MRI lymph node status was reported if the short-axis diameter of the largest reported lymph shown on MRI was equal to or greater than 1 cm. The hybrid model then outputs a predicted lymph node metastasis probability as an "H score."
| Performance of hybrid AI model for identifying lymph node metastasis on MRI in cervical cancer patients | ||
| Primary dataset | External test dataset | |
| Sensitivity | 93% | 90.6% |
| Specificity | 97.4% | 87.2% |
| Accuracy | 96.5% | 87.9% |
| Area under the curve | 0.963 | 0.933 |
What's more, the hybrid model's results were significantly associated with cervical cancer prognosis. Patients with higher H scores -- and therefore deemed to be high-risk -- had significantly shorter median disease-free survival time than those with lower scores. High-risk patients also had significantly lower probability of being disease-free (hazard ratio: 4.59, p < 0.001).
The researchers acknowledged the limitations of their study.
"First, a more extensive and prospective data set is needed to generalize the performance of the [deep-learning] model," they wrote. "Second, although [contrast-enhanced T1-weighted imaging] showed better performance than [T2-weighted imaging] and [apparent diffusion coefficient] maps, the combination of these sequences is unclear."