MRI software developer Random Walk Imaging is highlighting results from a study showing that its scanning method and software improved the sensitivity of MRI to detect white-matter brain changes related to multiple sclerosis (MS).
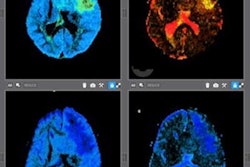
Random Walk's software uses an algorithm that converts standard fractional anisotropy (FA) into a microscopic fractional anisotropy (mFA) on diffusion-tensor MRI. The algorithm is designed to reduce the impact of white-matter fiber orientation on MRI sensitivity, the company noted.
In a study published June 8 in Brain Communications, researchers from the Danish Research Centre for Magnetic Resonance at Copenhagen University Hospital Hvidovre put the company's software to the test on patients with and without MS.
Random Walk's microscopic mapping algorithm improved the detail of MRI scans and helped clinicians to significantly improve the detection of disease-related cerebral white matter degeneration in patients with MS, the researchers found.
The reduction in mean mFA also had positive relationships with physical disability and total white-matter lesion load, as well as a positive correlation with individual cognitive dysfunction. In comparison, standard mean FA didn't identify relationships between normal-appearing white matter microstructure and clinical, cognitive, or structural measures, RWI noted.
The study authors concluded that mFA mapping "substantially advances the assessment of cerebral white matter in multiple sclerosis." Random Walk is currently conducting studies in Australia, China, and the U.S. to evaluate how its software can be clinically optimized for different anatomies.