It isn't every day that cheese is the focal point of a functional MRI (fMRI) study, but a foray by French researchers into an area of research that is seldom -- if ever -- explored earned them an Ig Nobel Prize.
Ig Nobel Prizes are bestowed by the Annals of Improbable Research, whose staff scours the globe for unique and quirky papers that may not have received the attention the authors so justifiably deserve.
Take, for example, the study of humans' aversion to some foods, such as cheese. In most scientific research, cheese would be fed to experimental mice.
In their prize-winning fMRI offering, however, Dr. Jean-Pierre Royet from Lyon Neuroscience Research Center and colleagues enrolled 15 people who like cheese and 15 people who do not care for fromage to analyze how their brains responded to its smell. Their work was published in Frontiers in Human Neuroscience on October 17, 2016.
"Although cheese is considered edible by most people, it can also be perceived as particularly disgusting to some individuals," the authors wrote. "As such, the perception of cheese constitutes a good model to study the cerebral processes of food disgust and aversion."
The participants were subjected to 40 different food fragrances, including six varieties of cheese (blue cheese, cheddar, goat cheese, Gruyère, Parmesan, and tomme) and nondairy items such as cucumber, fennel, mushroom, pâté, peanut, and pizza.
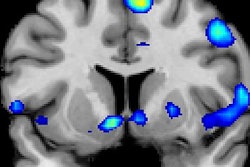
The individuals were imaged on a 1.5-teasla MRI scanner (Magnetom Sonata, Siemens Healthineers) and their brain activity on fMRI was recorded. Brain activity was based on three factors: a subject's interest in food and/or the cheese, whether the person liked or wanted the samples, and the person's reactions to the odors.
In the end, 37% of the subjects were "disgusted" by the smell of cheese. Approximately 60% of them objected to the odor and taste of cheese, while 18% reported a cheese intolerance or allergy.
For the record, the greatest brain activation related to liking and wanting cheese was seen in the globus pallidus. There was no such activation seen in the anticheese group.
"Food aversion has been underresearched because it is ethically inconceivable to experimentally induce illness in humans," the authors wrote. "In this study, we showed that a substantial proportion of people in France are disgusted by cheese and that this situation is experimentally favorable for studying the cerebral processes of food disgust and aversion."
There was no word from the researchers on which food group might be tested next.