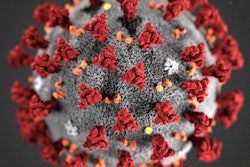
The SARS-CoV-2 virus that causes COVID-19 remains stable for several hours to days when it is aerosolized or on surfaces, according to a letter to the editor published online March 17 in the New England Journal of Medicine.
The virus was found in aerosols for up to three hours, on copper for up to four hours, on cardboard for up to 24 hours, and on plastic and stainless steel for up to two or three days, wrote researchers from the U.S. National Institutes of Health (NIH), the U.S. Centers for Disease Control and Prevention, the University of California at Los Angeles, and Princeton University.
"The results provide key information about the stability of SARS-CoV-2, which causes COVID-19 disease, and suggests that people may acquire the virus through the air and after touching contaminated objects," the team said in an NIH statement.
A group led by Neeltje van Doremalen, PhD, of the U.S. National Institute of Allergy and Infectious Diseases in Hamilton, MT, compared the stability of SARS-CoV-2 with SARS-CoV-1, which causes severe acute respiratory syndrome (SARS) (SARS infected more than 8,000 people around the world in 2002 and 2003 and is the virus most closely related to SARS-CoV-2). The two viruses have similar stability profiles, making it unclear why SARS-CoV-2 appears to be more infectious than its predecessor.
The authors highlighted the following observations:
- It appears that people infected with SARS-CoV-2 might be spreading the virus before they show symptoms, which makes disease control measures that were effective against SARS-CoV-1 less effective now.
- Unlike SARS-CoV-1, most secondary cases of SARS-CoV-2 transmission seem to be occurring in community rather than healthcare settings. But healthcare settings are vulnerable to the introduction and spread of SARS-CoV-2, and its stability in aerosols and on surfaces probably contributes to the transmission of the virus.
"The findings affirm the guidance from public health professionals to use precautions similar to those for influenza and other respiratory viruses to prevent the spread of SARS-CoV-2," the NIH said.