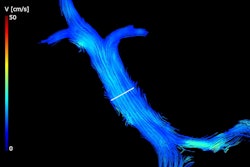
An MRI-based technique appears to be an effective, noninvasive option for diagnosing metabolic dysfunction-associated steatotic liver disease (MASLD) patients at risk of metabolic dysfunction-associated steatohepatitis (MASH), researchers have reported.
The findings address a clinical need, according to a group led by medical student Shyna Zhuoying Gunalan, of the National University of Singapore. Gunalan and colleagues published results from a literature review on January 13 in the American Journal of Gastroenterology.
"Non-invasive, accurate diagnostic tools are critical for identifying patients at risk [of MASH]," they noted.
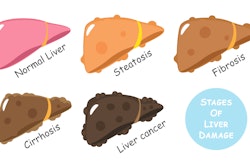
MASH is an advanced form of MASLD and is characterized by damage to the liver, inflammation, and varying degrees of fibrosis. Patients considered to be at risk have metabolic dysfunction-associated steatohepatitis activity scores (MAS) equal to or greater than 4, on a scale of 0 to 8.
Gunalan and colleagues conducted a search of Medline and Embase (from the databases' inception to December 2024), identifying studies that reported on MRI-based diagnostic techniques for "at risk" MASH. Twenty studies that included 9,480 participants met the team's inclusion criteria. The group calculated sensitivity, specificity, and diagnostic odds ratios, and applied "ruling in" (TRI) and "ruling out" (TRO) thresholds for two techniques: Focused Abbreviated Survey MRI (FAST) and MR elastography (MRE) combined with the Fibrosis-4 (FIB-4) Index for Liver Fibrosis (MEFIB).
The investigators reported the following:
- The FAST technique showed the highest TRO sensitivity (87%) with moderate specificity (57%) and TRI specificity of 90% with reduced sensitivity (44%).
- MEFIB showed high TRO sensitivity (81%) but lower specificity (60%). Its TRI specificity was 87%, and its sensitivity was 50%.
"FAST with its accessibility and robust diagnostic performance may be well-suited for large-scale application," the group concluded.
Access the full study here.