Using an artificial intelligence (AI) deep convolutional neural network (DCNN) to analyze x-ray images improves radiologist performance in identifying hip fractures, according to a study published July 23 in the European Journal of Radiology.
The combination could mitigate the at-times challenging task of spotting hip fractures on x-ray, wrote a team led by Dr. Tsubasa Mawatari of Teikyo University in Fukuoka, Japan.
The study suggests that "DCNN may have the potential to identify additional abstract features that have not been apparent to the human reader," the group wrote.
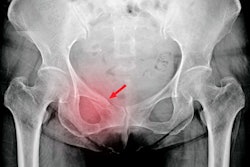
Hip fractures are a common cause of hospitalization in the elderly, and most are diagnosed with x-ray. But some patients have fractures that are hidden on these exams, perhaps due to overlying soft tissues or other technical factors. In fact, the rate of hidden hip fractures on x-ray is estimated to be between 3% and 10% of all negative hip or pelvic exams taken for trauma, according to the group.
In these cases, patients may have to undergo further imaging with CT or MRI to identify these fractures, which not only increases healthcare costs and exposes patients to further radiation but also causes treatment delays.
Mawatari and colleagues hypothesized that using a DCNN with x-ray to identify hip fractures could improve the interpretation performance of radiologists. They conducted a study that trained the AI algorithm using CT and MRI exams of patients with hip fractures rather than x-rays of hip fractures.
Seven radiologists read hip x-rays with and without the use of the AI algorithm. The study included 327 patients who underwent pelvic CT or MRI and were diagnosed with hip fractures; the AI algorithm for x-ray was trained with 302 of these exams, while the remaining 25 cases used for testing the algorithm. Mawatari's group measured radiologist and algorithm performance using the area under the receiver operating curve (AUC) measure.
The researchers found that the AI algorithm improved radiologist reader performance when it came to identifying hip fractures on x-ray.
| Reader performance for identifying hip fractures on x-ray, with AI and without | |||
| Readers alone | AI algorithm alone | Readers with AI | |
| AUC | 0.83 | 0.91 | 0.88 |
The study results show that AI offers a number of benefits for this particular clinical scenario, according to the authors.
A DCNN "has the potential to increase the efficiency of diagnosis and expand access to 'expert level' medical image interpretation," they concluded.