A 3D image-analysis software algorithm can identify small changes in the cartilage of knee joints over time on MRI exams, potentially paving the way for development of new osteoarthritis treatments, according to a study published May 24 in the Journal of Magnetic Resonance Imaging.
A team of researchers led by James MacKay, PhD, of the University of Cambridge in England developed 3D cartilage surface mapping (3DCaSM), a semiautomated image-analysis technique that creates a 3D model of the knee joint, as well as a "change map" that delineates cartilage changes between scans. In testing, the algorithm was able to detect significant changes in cartilage relaxation times and thickness over a period of six months that weren't detected using standard x-ray or MRI techniques.
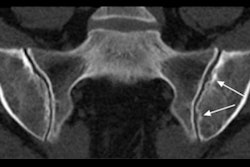
Osteoarthritis is traditionally identified on x-ray, but that modality doesn't offer enough sensitivity for detecting changes in the joint over time, according to the researchers. That makes it difficult to assess the effectiveness of treatments.
After previously developing an algorithm to monitor subtle changes in arthritic joints on CT scans, the group sought to apply these techniques to MRI in order to go beyond cartilage or bone thickness and gain more complete information about tissue composition. MRI is already used in clinical trials and observational studies to quantitatively assess arthritis, but manual segmentation of the data can be time-consuming, as well as less reproducible and sensitive to changes in the joint, according to MacKay and colleagues.
To address this limitation, the team developed 3DCaSM to analyze cartilage relaxation times and morphology on contrast-enhanced MRI. The algorithm utilizes modified cortical bone mapping and joint space mapping techniques, according to the researchers.
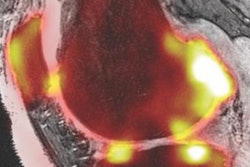
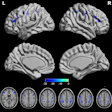
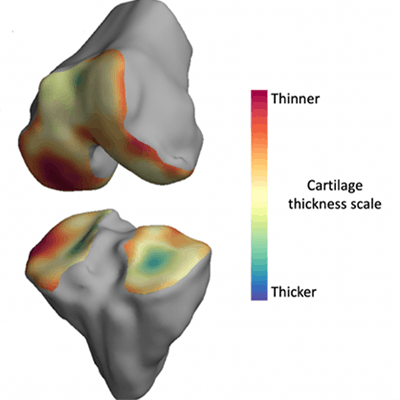
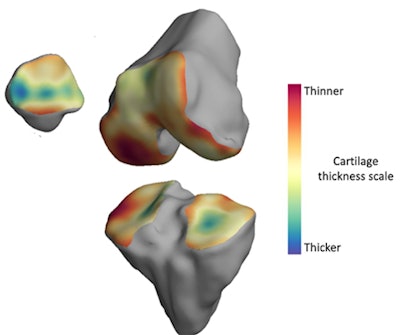 The thickness of the cartilage covering the end of each bone is color-coded, with red areas denoting thinner cartilage and green-blue areas denoting thicker cartilage. The technique helps locate where arthritis is affecting the joint over time. Image courtesy of the University of Cambridge.
The thickness of the cartilage covering the end of each bone is color-coded, with red areas denoting thinner cartilage and green-blue areas denoting thicker cartilage. The technique helps locate where arthritis is affecting the joint over time. Image courtesy of the University of Cambridge.The researchers validated the algorithm on four cadaver knees, then tested it on 14 study participants between the ages of 40 and 60 with mild to moderate knee osteoarthritis. The software was also used to assess six study participants in the same age range with no joint pain at baseline, one month, and six months.
The algorithm showed significant six-month changes in study participants' cartilage thickness and relaxation times in both those with osteoarthritis and those without the disease. The MRI parameter with the greatest change at six months was T2 relaxation time, according to the authors.
"There's a certain degree of deterioration of the joint that happens as a normal part of aging, but we wanted to make sure that the changes we were detecting were caused by arthritis," MacKay said in a statement from the university. "The increased sensitivity that 3D-CaSM provides allows us to make this distinction, which we hope will make it a valuable tool for testing the effectiveness of new therapies."
The algorithm's findings highlight the spatially heterogeneous and bidirectional nature of these cartilage changes, according to the authors.
"Our results suggest that the use of 3D‐CaSM and similar pipelines is likely to offer improved responsiveness when compared to approaches that fail to recognize this heterogeneity; for example, approaches that involve analysis of only a single subregion and assume unidirectional change," they wrote.
The software is free to download and can be added to existing systems, according to the researchers. The authors also noted that they will be assessing the algorithm's ability to predict a patient's need for a knee replacement as part of a new study funded by the European Union.