A newly developed deep-learning artificial intelligence (AI) model is lending a helping hand to radiologists reading functional MRI (fMRI) scans, helping them better predict attention deficit hyperactivity disorder (ADHD), according to a study published online December 11 in Radiology: Artificial Intelligence.
The multichannel deep neural network (mcDNN) model created by researchers from the University of Cincinnati College of Medicine and Cincinnati Children's Hospital Medical Center used a combination of multiscale brain connectome data and personal patient information to achieve better accuracy in ADHD detection than a single-channel deep neural network with the same input.
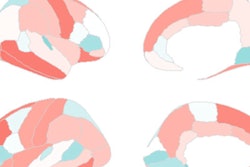
"Our results emphasize the predictive power of the brain connectome," said senior author Lili He, PhD, from Cincinnati Children's Hospital Medical Center, in a statement. "The constructed brain functional connectome that spans multiple scales provides supplementary information for the depicting of networks across the entire brain."
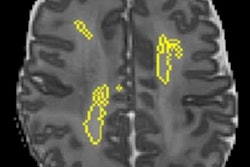
Recent advances in fMRI techniques have allowed researchers to quantitatively map connections within and between brain networks. This functional connectome is critical to understanding the specific etiology behind many brain disorders, including ADHD, and is constructed from spatial regions across the fMR images known as parcellations. The brain parcellations are based on anatomical criteria, functional criteria, or both, and allow the brain to be studied at different scales based on different brain parcellations.
Concurrently, deep-learning neural network models have begun to supersede traditional machine-learning algorithms to better classify and characterize brain disorders by producing "physiologically meaningful features" and revealing "new associations from high-dimensional neuroimaging data," the authors wrote. "To this end, we proposed a multichannel deep neural network (mcDNN) model analyzing multiscale functional connectome data and tested the value of this model by using ADHD detection as an example."
For this retrospective study, He and colleagues collected data from 973 participants in the Neuro Bureau ADHD-200 repository in order to develop their mcDNN model from multiscale functional brain connectomes based on the subjects' personal anatomic and functional data. Their brain connectomes featured one anatomically defined template with 90 parcellated regions of interest (ROIs) and two functionally defined templates with 190 parcellated ROIs and 351 parcellated ROIs, respectively.
The researchers then compared their mcDNN model with a single-channel deep neural network (scDNN) model for sensitivity, specificity, accuracy, and area under the receiver operating characteristic curve (AUC).
The mcDNN model significantly outperformed the scDNN with an accuracy of 78%, compared with the scDNN, which had its greatest accuracy of 73% when only personal patient data was included in the model. In addition, the mcDNN registered a statistically significant difference in AUC at 0.82, compared with an AUC of 0.67 for the scDNN (P < 0.001).
Given the results, the researchers believe using deep-learning models to aid MRI-based diagnosis could be critical in implementing early interventions for ADHD patients and could be valuable with other neurological disorders.
"We already use it to predict cognitive deficiency in preterm infants," He added. "We scan them soon after birth to predict neurodevelopmental outcomes at two years of age."
In fact, the researchers plan to bring this deep-learning model to analyze larger neuroimaging datasets, with a goal to better detect and understand dysfunctions in the connectome and their association with brain disorders.