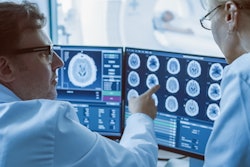
Accelerating MRI protocols can reduce greenhouse gas emissions, energy use, and costs, while also improving patient access and revenue, according to a study published April 22 in Radiology.
The finding is from a retrospective analysis that used MRI clinical and phantom acceleration data, with protocol acceleration showing a linear relationship between time and energy reduction, noted lead author Sean Woolen, MD, of the University of California, San Francisco, and colleagues.
“These findings highlight the value of sustainable operations and energy-saving strategies,” the group wrote.
The high electricity demand of MRI makes it a key target for energy-saving strategies, and although nonoperational reductions have been studied, data on the impact of scan time reduction techniques are limited, the authors explained.
To address the knowledge gap, the group first gathered data logs from 14 days from three Siemens Healthineers MRI scanners: Magnetom Vida 3T, Magnetom Sola 1.5T, and 0.55-T Magnetom Free.Max 0.55T. From the data, they calculated total and sequence-specific energy consumption, examination duration, and carbon emissions for 377 clinical examinations.
Next, the team used standardized phantom experiments to test three acceleration techniques: multiband simultaneous multislice (SMS, Siemens Healthineers), parallel acceleration (PAT, syngo Grappa, Caipirinha, Siemens Healthineers), and deep learning (DL)-based reconstruction (Deep Resolve; Siemens Healthineers).
Acceleration settings included PAT with acceleration factors of 2 and 3, SMS with an acceleration factor of 2 combined with PAT-2 (total factor of 4), and DL with SMS-2 and PAT-2 (total factor of 4).
A comparative analysis showed a linear relationship between protocol acceleration and time and energy reduction, according to the findings. Accelerating sequences by 25% to 75% lowered per-examination greenhouse gases and energy by 21% to 65%, with deep learning having the greatest impact.
The researchers calculated that for a seven-day practice with 24 MRI examinations, projected annual savings from the accelerated protocols ranged from 200.6 to 867.1 MWh, $26,073 to $112,719, and 140.2 to 606.1 metric tons of carbon dioxide equivalent (CO2eq). In addition, the techniques were estimated to enable from 8,424 to 57,564 additional appointments and thus generate between $3.3 million and $22.2 million in revenue.
“DL and other acceleration methods cut [greenhouse gas] emissions, energy use, and costs while boosting MRI capacity and revenue,” the group wrote.
The authors noted that the lower-field-strength scanners used less energy, but increased examination times, with phantom experiments showing significant differences only between field strengths of 3-tesla and 0.55-tesla under matched protocols.
Limitations included reliance on three MRI scanners from one vendor, modeling from clinical and phantom data, variability in energy estimates across institutions, exclusion of chilled water energy, and the use of U.S. national averages for energy costs, reimbursement, and carbon savings, the group added.
Nonetheless, the study significantly demonstrated that protocol modifications can reduce the environmental impact of MRI, they wrote.
“Optimizing MRI protocols with acceleration methods such as DL offers substantial environmental and operational benefits, reducing [greenhouse gases], energy use, and costs while improving patient access and revenue,” the researchers concluded.
Study authors included employees of Siemens Healthineers, with non-industry authors maintaining full control of data and analysis, the group noted.
The full study is available here.