A deep-learning model performs comparably to an abdominal radiologist when it comes to finding clinically significant prostate cancer on MRI, researchers have reported.
The model shows promise as a way to improve radiologists' diagnostic performance on MRI by increasing cancer detection rates and decreasing false positives, study lead author Naoki Takahashi, MD, from the Mayo Clinic in Rochester, MN, said in a statement released by the RSNA. The study findings were published August 6 in Radiology.
"[It's not that] we can use this model as a standalone diagnostic tool," he said, "[but] the model's prediction can be used as an adjunct in our decision-making process."
Prostate cancer is the second most common in men around the world, and radiologists use MRI to diagnose clinically significant disease; findings are expressed through the Prostate Imaging-Reporting and Data System version 2.1 (PI-RADS). But classifying prostate lesions with PI-RADS can be tricky -- which is why using AI algorithms with prostate MRI could improve cancer detection and reduce observer variability, the group explained.
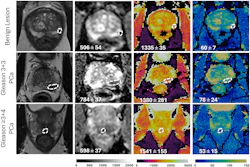
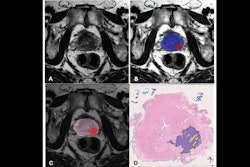
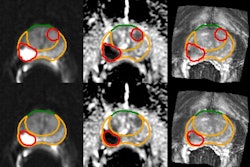
Takahashi's team trained a convolutional neural network to predict the presence of clinically significant prostate cancer on multiparametric MR imaging (T2-weighted, diffusion-weighted, and T1-weighted contrast-enhanced images, as well as apparent diffusion coefficient maps). The group compared the model's performance to that of abdominal radiologists via a study that included 5,215 patients without known clinically significant prostate cancer who underwent MRI between January 2017 and December 2019. Of the total study cohort, 1,514 examinations showed clinically significant prostate cancer; to train the model, the investigators used 400 exams as an internal test set and 204 as an external test set.
Overall, the model performed comparably to experienced abdominal imaging radiologist readers for identifying prostate cancer, and a combination of the model and the reading radiologist findings performed better than radiologists alone on both internal and external test sets (area under the receiver operating curve [AUC] of 0.89 (p < 0.001).
| AUC of deep-learning model compared with radiologist readings for identifying clinically significant prostate cancer | ||
|---|---|---|
| Test set type | Deep-learning model | Radiologist reader |
| Internal | 0.89 | 0.89 |
| External | 0.86 | 0.84 |
The study highlights the potential of integrating deep learning with MRI to address "long-standing challenges in [prostate cancer] diagnosis," noted Patricia Johnson, PhD, and Hersh Chandarana, MD, both of NYU Langone Health in New York City, in a commentary.
"As we continue to refine these technologies and methods, the goal of providing effective, accessible, and cost-efficient screening tools moves closer to reality," Johnson and Chandarana wrote. "This progress promises to improve patient outcomes through earlier and more accurate diagnosis of [clinically significant prostate cancer]."
The complete study can be found here.