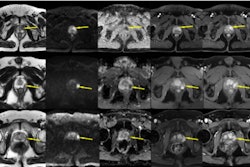
It's better to take Prostate Imaging-Reporting and Data System (PI-RADS) upgrading rules into account when it comes to deciding whether a patient with clinically significant prostate cancer needs a biopsy, researchers have reported.
"[Our] study supports the application of PI-RADS upgrading rules to optimize biopsy selection, particularly in risk-based pathways," wrote a team led by Eduardo Correia, MD, PhD, of University Hospitals in Cleveland, OH. The study results were published February 14 in the American Journal of Roentgenology.
PI-RADS incorporates rules by which ancillary findings can upgrade a dominant score to a higher final category, the group noted. But evidence as to how this type of upgrade impacts various pathways for diagnosing prostate cancer is unclear. To address the knowledge gap, the team evaluated the benefit of using PI-RADS upgrading rules with MRI-directed diagnostic sequences.
Correia and colleagues conducted research that included 716 patients without known prostate cancer who underwent prostate MRI followed by biopsy between January 2016 and May 2020. The investigators defined clinically significant prostate cancer as being in a grade group equal to or greater than 2, and they compared the detection of disease between dominant (that is, no upgrade rule applied) and upgraded lesions.
They also assessed net benefit of detecting clinically significant prostate cancer and avoiding biopsy using the following MRI-directed pathways (which both considered and disregarded PI-RADS upgrading rules):
- A biopsy-all pathway
- An MRI-focused pathway (i.e., no biopsy for PI-RADS equal to or less than 2)
- A risk-based pathway (i.e., biopsy in patients with a prostate-specific antigen density equal to or more than 0.15 and a PI-RADS score equal to or less than 3)
The authors reported no significant differences for diagnosis of clinically significant prostate cancer between dominant and upgraded PI-RADS 3 transition zone lesions (20% compared with 19%), PI-RADS 4 transition zone lesions (33% compared with 26%), or PI-RADS 4 peripheral zone lesions (58% compared with 45%). They also found the following:
| Frequencies of clinically significant prostate cancer by diagnostic pathway | |||||
|---|---|---|---|---|---|
| Measure | Biopsy-all | MRI-focused | MRI-focused without regard for upgrading rules | Per-guideline risk-based | Risk-based without regard for upgrading rules |
| Frequency | 53% | 52% | 51% | 52% | 48% |
| Biopsy avoidance | 0% | 13% | 16% | 19% | 25% |
"Disregarding PI-RADS upgrading rules reduced net clinical benefit of the risk-based MRI-directed diagnostic pathway when considering tradeoffs between clinically significant prostate cancer detection and biopsy avoidance," the authors concluded.
The complete study can be found here.