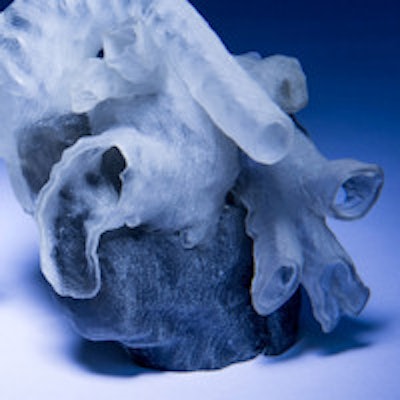
Researchers from the Massachusetts Institute of Technology (MIT) and Boston Children's Hospital have developed an automated software method for converting 3D MRI scans of a pediatric patient's heart into a tangible model that surgeons can use to plan operations.
While the endeavor to create heart models from 3D MRI scans is not unique -- other institutions around the country are involved in similar projects -- this initiative is looking to craft a faster and more automated process to define the borders and contours of pediatric hearts and facilitate the production of 3D models.
 Dr. Andrew Powell, senior associate in cardiology at Boston Children's Hospital.
Dr. Andrew Powell, senior associate in cardiology at Boston Children's Hospital."Each patient's heart is a little bit different," said Dr. Andrew Powell, a senior associate in cardiology at Boston Children's Hospital. "That puts a big premium on accurate information, because [cases] do not fall into just two or three different categories."
How to build a heart
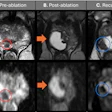
Typically, clinicians and cardiologists use echocardiography or angiography to obtain 2D images of the abnormal areas of a child's heart. Physicians then use the information to try to visualize a "three-dimensional picture of the problem," Powell said.
"It would be better and probably more useful to have a 3D representation of that particular heart anatomy, rather than having to rely on a 2D integration in people's heads," he added.
The model-making process begins by having a clinician label and manually segment 10 to 15 short-axis slices of a 3D MR image.
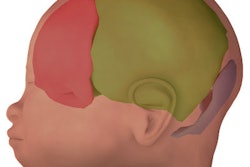
"We want to label the blood pool inside the heart chamber; the myocardium, including the walls around the atria and great vessels; and the background pixels, which would include, for example, the liver and the chest wall," said Danielle Pace, a graduate student in electrical engineering and computer science who is heading the development of the software to analyze the MRI scans.
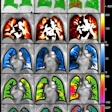
The researchers are using an algorithm known as patch-based segmentation to examine patches around each unlabeled voxel and find the most similar patches in the labeled examples. By doing so, the algorithm can achieve 90% agreement with expert segmentation if 14 short-axis slices are manually labeled, and 80% agreement if three short-axis slices are manually labeled.
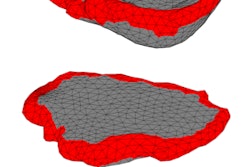
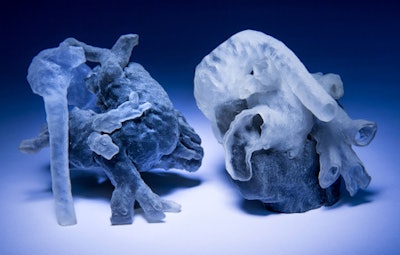 3D-printed heart models created from MRI scans. Image courtesy of MIT and Boston Children's Hospital. Photo credit: Bryce Vickmark.
3D-printed heart models created from MRI scans. Image courtesy of MIT and Boston Children's Hospital. Photo credit: Bryce Vickmark."The segmentation challenge was very interesting because it is still very difficult for image segmentation algorithms to handle cases with very severe anatomical variability or severe pathologies," Pace said.
To the best of her knowledge, this venture is the first time that patch-based segmentation has been used as an interactive technique, Pace said.
"It is usually used to propagate data across subjects," she added. "This time we are doing it within the same subject, in this case for whole-heart segmentation in congenital heart disease."
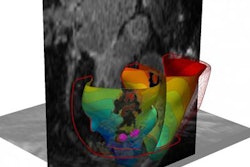
After the labeling information is obtained, the researchers use a marching cubes computer graphics algorithm to convert the segmentation information into a tangible model that can be viewed on screen or 3D-printed.
 Danielle Pace, MIT graduate student in electrical engineering and computer science.
Danielle Pace, MIT graduate student in electrical engineering and computer science."Because each heart is a little bit different and a lot of those borders need to be drawn manually, what we are looking to do is to come up with a faster, more automated, less labor-intensive way of finding those heart borders to facilitate the production of 3D heart models," Powell said. "Once we have those borders, the third step is printing the actual model. Each step of the pipeline needs to be efficient as well as practical."
Clinical evaluation
The 3D models have yet to be used in the clinical setting, but about a half-dozen cardiac surgeons at Boston Children's Hospital plan to collaborate on a study within the next two years to evaluate the models' utility. Powell will lead the research, though he will not be directly involved in the evaluations.
The study will take a retrospective approach, looking at past cases to see if 3D models would help with surgical planning.
"We have some cases from before, where we know what the surgical decisions were," Pace said. "We want to present the surgeons with the conventional MRI view, and/or a preprinted model, and/or a virtual model on the computer."
Essentially, the surgeons would plan surgery using these different scenarios, with the goal of determining if they would make the same decisions. One question is whether surgeons would prefer a larger model to see imperfections of the heart more precisely, or would they prefer a smaller, life-sized model to reflect the close confines in which they need to operate.
"A true-to-size model has an advantage because these models could be used to practice the procedure," Powell said. "But for just visualization and teaching, it might be better to have something that is a little larger and easier to work with."
Pace and Powell, along with colleagues Polina Golland, PhD, a professor of electrical engineering and computer science at MIT, and Mehdi Moghari, PhD, a physicist at Boston Children's Hospital, plan to present their new system at the International Conference on Medical Image Computing and Computer Assisted Intervention (MICCAI) in October in Munich.
The project was funded by Boston Children's Hospital and by Harvard Catalyst, a consortium designed to support scientific research and bring new technologies into clinical settings.