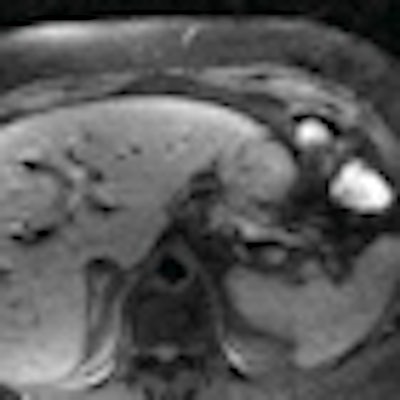
Understanding how contrast-enhanced MRI underestimates tumor size compared to pathologic specimens may allow more precise assessment of tumor volume prior to radiation therapy for pancreatic cancer, according to researchers from Emory University.
The group from Emory's Winship Cancer Institute found that the accuracy of tumor size by contrast-enhanced ultrafast gradient echo (3D VIBE) MRI is critical, but they warned that MRI becomes less accurate as the tumor grows larger.
"The potential benefits of using MRI for treatment planning arise from more accurate definition of target volume," said lead study author Dr. John Mikell, a resident in the department of radiation oncology. "Better target delineation may translate into less tumor missed and less normal tissue treated, which may in turn translate into better tumor control and decreased toxicity."
Mikell presented study results at the American Society for Radiation Oncology (ASTRO) annual meeting in Boston.
MRI vs. CT
Contrast-enhanced CT is currently the standard choice for tumor delineation because CT images are most commonly obtained preoperatively, according to the researchers. However, CT also underestimates tumor size compared to pathological results.
MRI, Mikell noted, provides a high level of tumor conspicuity for pancreatic cancers, which makes the modality "highly valuable" for delineating target volumes in radiotherapy planning without exposing a patient to radiation from a CT scan.
"Although determination of resectability is very important for the patient's initial staging, radiation oncologists are more concerned with conspicuity, as we need to accurately define a tumor for treatment planning," Mikell explained.
Accurate target delineation is increasingly important as conformal, intensified radiotherapy techniques, such as stereotactic body radiation therapy (SBRT) and intensity-modulated radiation therapy (IMRT), are associated with better local tumor control, he added.
For this study, Mikell and colleagues reviewed records of 209 patients (median age, 66 years) who underwent pancreatic cancer resection between 2000 and 2011. Of those patients, 92 had both preoperative contrast-enhanced 3D VIBE MRI and detailed specimen measurements available for evaluation.
Primary tumor size from the MR images was measured independently by a diagnostic radiologist blinded to pathology reports. Pathology tumor measurements from gross specimens were obtained from the patients' pathology reports. The researchers then compared the maximum dimensions of tumors on MRI with the results from the gross specimens. The median difference between the pathology samples and MRI measurements was calculated.
Results
The pancreatic head was the primary site among 80 patients (87%) and body or tail was the primary site in 13 (14%). Three patients were confirmed as having stage IA pancreatic cancer, while seven had stage IB cancer. In addition, 21 patients were determined to have stage IIA pancreatic cancer, 57 had stage IIB cancer, and four had stage III disease.
A review of the data found that MRI underestimated tumor size by a median of 4 mm (range, -22 to +34 mm). The median largest tumor dimension on MRI was 2.65 cm (range, 1.5 to 9.5 cm), compared with a median of 3.2 cm (range, 1 to 10 cm) for the pathology specimens. On average, the pathology specimens were 5.1 mm larger than the largest MRI measurements.
Based on the results, Mikell and colleagues concluded that MRI could be used in conjunction with CT for radiation planning, as most radiotherapy treatments are based on a CT simulation scan.
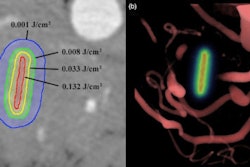
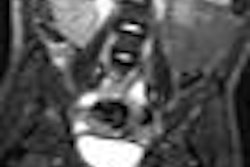
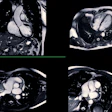
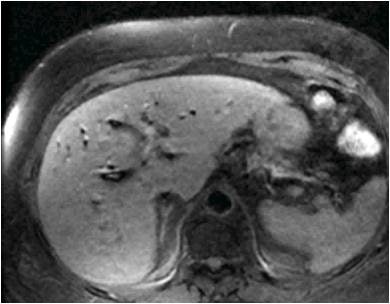 |
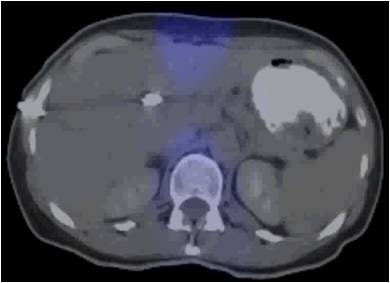
| The 3D VIBE MR image (above) shows the difference in data acquired compared to CT with stereotactic body radiation therapy (below). All images courtesy of Dr. John Mikell. |
 |
At Emory, the radiotherapy plan is calculated using the CT simulation, which is linked to the center's 3D coordinate system and radiation therapy treatment machines. "During treatment planning, however, MRI may be fused with the CT using registration software, which allows the practitioner to use the MR images to help define the target volume," Mikell said. "This is the current practice at our institution."
Even though the discrepancy for MR images is slightly less than when CT results are compared to pathology specimens, the authors noted that novel imaging techniques are needed to more accurately delineate tumors.
Mikell and colleagues plan to continue their research by investigating the radiographic and pathologic correlation in pancreatic cancer using a variety of imaging modalities, all with the goal of improving target volume delineation.