The image interpretation process has come a long way since the days of static film. In the next evolutionary phase, radiologists may be given the ability to perform interactive reconstruction of raw CT projection data -- the sinogram -- according to Dr. Eliot Siegel of the University of Maryland.
In addition to being able to zoom in on a particular section of an image to access the full sampled data in the sinogram, this would allow radiologists to deploy particular reconstruction kernels based on specific examination type or exam indication. They could also optimize their evaluation by being able to apply multiple types of processing techniques dynamically and in real-time on different areas of the same image, Siegel said.
"I believe there may be advantages to this approach that could lead to improved diagnostic efficacy and confidence," he said.
Although access to sinogram data is typically not provided by vendors, Siegel and colleagues are collaborating with two firms to develop a method for facilitating on-demand reconstruction of sinogram data.
"Reconstruction on demand [could] provide the opportunity for innovation in the continuing evolution of image visualization at the workstation," Siegel said.
He discussed the potential for bringing advanced reconstruction of sinogram data to the workstation during a session at the recent International Symposium on Multidetector-Row CT in Washington, DC.
What is raw data?
Reconstruction of raw image data is a frequent topic in CT, but there isn't an exact definition of what "raw data" represents, Siegel said.
"Is it the CT images? Is it the thin-section data?" he asked. "One thing I've been interested in for quite a while is looking at the potential advantages of being able to actually interact at the workstation with the true raw data, which is the sinogram data."
The first phase of the radiologist image interpretation process was the film era. In the second phase, radiologists transitioned to PACS and soft-copy interpretation in static "tile" mode similar to film, Siegel said.
Dynamic image interaction arrived in the third phase, as radiologists learned to rapidly -- and, eventually, unconsciously -- window and level and use other workstation tools. The next phase, stack mode and linked stack mode, took advantage of the visual system's enhanced ability to detect motion or other changes in a visual field, according to Siegel.
"As we go from the passive, film-based type of mentality to a more interactive mentality, the radiologists, technologists, and clinicians played a much more active role in interacting with the datasets," he said.
We've now reached the fifth phase, which makes use of volumetric and multiplanar navigation, Siegel said.
"We have the capability of thinking of the image set as a volume," he said. "But that volumetric image set is actually a series of axial, thin-section data that we can interact with now that it's isotropic."
The future
The sixth phase may actually support interactive review of the CT sinogram data itself, Siegel said. This would require ultrahigh-speed processing and the capability to zoom in to the full sinogram data, as well as the ability to change on the fly the kernel of reconstruction for an image, or at least a portion of the image.
"Wouldn't it be great to identify a portion of the image and be able to zoom in on it?" he asked.
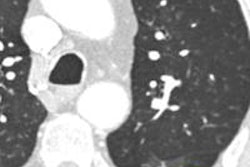
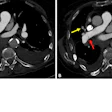
 The ability for radiologists to interact directly with the raw CT data (sinogram) may lead to improved diagnostic efficacy and confidence. Image courtesy of Dr. Eliot Siegel.
The ability for radiologists to interact directly with the raw CT data (sinogram) may lead to improved diagnostic efficacy and confidence. Image courtesy of Dr. Eliot Siegel.
This evolution would be analogous to what took place in digital radiography. Initially, radiologists sought to have images that reproduced the look of film as faithfully as possible, and workstation tools that mimicked traditional film interpretation processes. However, those preferences changed over time.
"Vendors started learning through customer feedback that they could process the images and enhance the images, potentially interactively with a radiologist," Siegel said. "And one of the things we found with digital x-ray was that there was no one ideal mechanism or look or feel for the images."
What worked well for the lung didn't work well for the mediastinum, for example, while what worked well for the mediastinum might not work for the bones or the abdomen.
"In an analogous way, is it not possible that in CT, there might be specific mechanisms or ways to be able to do the processing, such as applying iterative reconstruction or different kernels, that would be optimized and have multiple different settings in the same way we do window and level settings?" he suggested.
All exams and patients are not alike, and several factors must be accounted for, including anatomic region, patient body habitus, technique, and the specific type of pathology being evaluated, he said. In digital x-ray, a number of different processing techniques have been developed to see more information and to be able to optimize the amount of data seen on a single image.
Is one image enough?
These dynamics raise the question of whether one image is enough for CT.
"Is there an advantage to multiple presentations of one image, and can we, for example, have specific processing for different disease states, as we've suggested for digital x-ray?" he asked. "It also raises the question of whether a single processing technique or algorithm is OK for CT, or if it's just a compromise."
Radiologists currently have the ability to select a reconstruction kernel based on a specific examination type or even exam indication, such as for interstitial lung disease, he said. However, on the same examination, they might also want to optimize evaluation of the mediastinum, searching for a liver lesion, or evaluation of the renal arteries, by being able to apply multiple types of processing dynamically and in real-time, Siegel said.
"What if we could go beyond our current limitation right now, which is basically window/level adjustment when we get the images?" he asked. "What if we could find a specific lesion and then be able to interrogate the sinogram data associated with the lesion?"
Currently, access to sinogram data isn't typically available, as sinogram data and reconstructions are considered by CT vendors to be their "special sauce," according to Siegel.
"The model that we have is that sinogram data gets reconstructed into CT sections and then we interact [using] our advanced workstations or PACS workstations with those derivative images," he said. "And vendors have understandably been unenthusiastic about sharing their sinogram data, except for research purposes. This accounts to some extent for a lot of the variability that we who do clinical trials and try to do quantitative measurements see from one scanner to another."
An alternative method
The University of Maryland is currently working with Hitachi Medical Systems and a third-party advanced visualization workstation (TeraRecon) to develop an alternative that could facilitate interaction with the sinogram data at the workstation.
Getting around the problem of gaining direct access to the sinogram data, this work-in-progress method would provide bidirectional communication between a PACS or advanced visualization workstation and the CT reconstruction engine. It would also provide zoom capability and on-the-fly image reconstruction for either the entire set or subset of the sinogram image dataset.
 Dr. Eliot Siegel from the University of Maryland.
Dr. Eliot Siegel from the University of Maryland.
"I would be able to dynamically change slice thickness and orientation, be able to turn on and off artifact correction for metallic artifacts or bowel gas, be able to select different iterative reconstruction parameters in a dynamic fashion, and perform the type of study that I want," Siegel said.
The prototype is continuing to be refined, and Sigel anticipates that testing will begin immediately once the institution takes delivery of Hitachi's 128-slice CT scanner in a few months.
"We will use a 'dedicated' TeraRecon server and will access it using a workstation that we will use for primary image interpretation of CT studies," Siegel told AuntMinnie.com. "We will test not only the clinical efficacy, but also the performance of the system to allow interaction between the radiologist and the sinogram data by having the TeraRecon server initiate requests to the Hitachi reconstruction engine to perform image reconstruction on the raw data (sinogram)."
On-demand reconstruction provides a number of advantages, he noted.
"Since most CT scanners don't currently support a 768 x 768 matrix size or greater, for example, we have the ability to subset, on the fly and on demand, a portion of the image and be able to get the full sampled data that's in the sinogram data," he said.
It also offers the potential to utilize parameters for specific tasks, such as measurement or characterization of a nodule or mass, and to optimize visualization for specific tasks, he said. Parameters such as artifact reduction could be applied on demand without altering the rest of the image.
Challenges to overcome
There are significant challenges to on-demand reconstruction, however. The volume and complexity of CT data are already high, and it's not intuitive whether or not this would add additional image interpretation time, he said.
In addition, reconstruction speeds are relatively slow, especially with iterative reconstruction, so it needs to be determined how to make this work rapidly and dynamically with today's technology, Siegel said. Furthermore, no standard exists for workstation interaction with the CT reconstruction engine.
That said, there's a major opportunity for interesting clinical research to evaluate the possible added clinical value of being able to interact dynamically with the sinogram dataset, Siegel concluded.