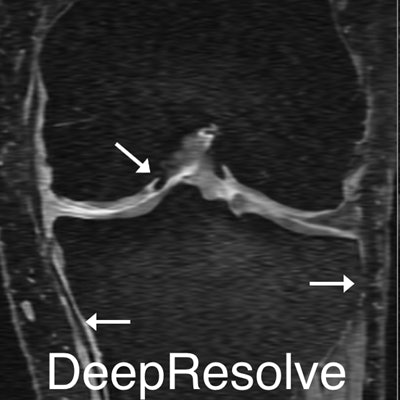
A deep-learning algorithm can transform low-resolution, thick-slice knee MR images into high-resolution, thin-slice images -- potentially yielding faster image acquisition, lower costs, and fewer motion artifacts in musculoskeletal MRI, according to research published online in Magnetic Resonance in Medicine.
In a pilot study, researchers from Stanford University trained a 3D convolutional neural network (CNN) to generate simulated high-resolution, thin-slice images from slices acquired at one to eight times greater thickness. In testing, their model -- called DeepResolve -- was both quantitatively and qualitatively superior to commonly used resolution-enhancement methods, according to first author Akshay Chaudhari, PhD, a postdoctoral research fellow.
"Superresolution methods such as DeepResolve can have an immediate impact in clinical studies as well as research studies," Chaudhari told AuntMinnie.com.
State of MSK imaging
The current state of clinical and research musculoskeletal imaging was the primary motivation behind the research project, Chaudhari said. Clinical MSK imaging typically involves 2D fast spin-echo sequences that provide high in-plane resolution but very thick slices and often also slice gaps.
 Akshay Chaudhari, PhD, from Stanford University.
Akshay Chaudhari, PhD, from Stanford University."Due to this, partial volume effects are common and protocols are often repeated with the same contrasts in different scan planes," Chaudhari said. "This reduces the efficiency of imaging, which is becoming more and more important as we move toward value-based imaging."
From a research point of view, there's a need to quantify subtle changes in tissue such as the cartilage and menisci to assess the progression of osteoarthritis, Chaudhari said. In these cases, it's challenging to balance the need for high resolution in segmenting tissues with the requirement of a high signal-to-noise (S/N) ratio to perform quantitative imaging.
"Thin slices are ideal for morphological assessment, [but] their low S/N ratio leads to inaccurate quantitative measurements," he said. "Our goal with this study was to be able to enhance through-plane resolution without compromising in-plane resolution to make MSK scans more efficient and to overcome S/N ratio limitations for quantitative imaging," he said.
The researchers trained DeepResolve with unsupervised learning on 124 double echo steady-state (DESS) knee datasets from the Osteoarthritis Initiative. All images had a slice thickness of 0.7 mm. They then tested the algorithm on 17 datasets (Magn Reson Med, March 29, 2018).
Quantitatively better
In quantitative testing, DeepResolve yielded significantly higher (p < 0.05) structural similarity, peak S/N ratio, and root-mean-square error than the commonly utilized Fourier interpolation, tricubic interpolation, and sparse-coding superresolution methods for nearly all downsampling factors, according to the researchers.
Two board-certified MSK radiologists then qualitatively ranked ground-truth images, DeepResolve images produced from images that had three times the thickness, and tricubic interpolation images. After reviewing the diagnostic quality of the three image sets for sharpness, contrast, artifacts, S/N ratio, and overall diagnostic quality, the radiologists preferred the DeepResolve images to those generated by the tricubic interpolation method commonly used in PACS and DICOM viewers, Chaudhari said. DeepResolve had significantly higher scores (p < 0.01) than the tricubic interpolation images in all image quality categories as well as overall image ranking.
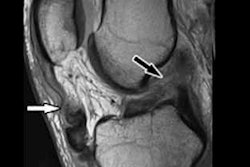
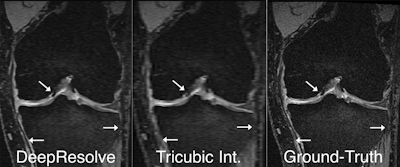 DeepResolve was found to enhance the image quality of blurry input images compared with other methods such as tricubic interpolation, which is commonly used in PACS and DICOM viewers. Example images demonstrate the inherent blurring present in coronal and axial reformations of sagittal acquisitions and how DeepResolve can eliminate much of the multiplanar reformation blurring. Images courtesy of Akshay Chaudhari, PhD.
DeepResolve was found to enhance the image quality of blurry input images compared with other methods such as tricubic interpolation, which is commonly used in PACS and DICOM viewers. Example images demonstrate the inherent blurring present in coronal and axial reformations of sagittal acquisitions and how DeepResolve can eliminate much of the multiplanar reformation blurring. Images courtesy of Akshay Chaudhari, PhD.Chaudhari and colleagues also concluded that DeepResolve did not seem to affect the visualization of pathology.
While the image quality produced by DeepResolve did not match the ground-truth images, the researchers noted that the ground-truth dataset isn't entirely clinically feasible; acquisition of that sequence took approximately 11 minutes, which is almost as long as an entire clinical imaging knee protocol. They intentionally chose that dataset for training the algorithm because it offered much higher-resolution images than standard clinical protocols, according to Chaudhari. The images in the dataset had a slice thickness of 0.7 mm, while clinical protocols use a slice thickness between 2.5 mm and 3 mm.
"By training our network on a standard that is better than the clinical protocol, our goal is to be able to enhance the clinical protocol itself," he said.
The algorithm requires only 10 seconds of inference time to produce its high-resolution images from a large dataset with 160 slices, according to the researchers.
Improving clinical practice
DeepResolve could improve clinical practice in a couple of ways, Chaudhari said. Because the method offers a mechanism for acquiring low-resolution slices and transforming them into higher resolution, scan times could be reduced for acquiring higher-resolution data. In addition, the method could enable multiplanar reformatting and alleviate the need for scanning with sequences that offer the same exact contrast but in a different scan plane.
"Both such methods can directly lower the scan duration in examinations, which is a big push in value-based clinical practice," he said.
While the research results were promising, Chaudhari noted that caution should be exercised in evaluating this technology. In addition, further studies should explore any systematic biases as well as the generalizability of these superresolution methods.
"There is a need for performing systemic studies in evaluating the impact of DeepResolve on pathologies," Chaudhari said.
The researchers are now actively trying to improve the performance of DeepResolve. While they have filed for a provisional patent, that doesn't preclude them from sharing and collaborating with noncommercial entities, he said.
"At the end of the day, methods like DeepResolve can only succeed if truly evaluated by the entire community," Chaudhari said. "That is why we want to share our research, code, and learned weights with the community so that we can collectively learn the strengths and weaknesses of the method and hopefully improve them to have a tangible impact."
Chaudhari and colleagues also presented a poster on this research at NVIDIA's recent GPU Technology Conference in San Jose, CA. It was named the best healthcare poster and best overall poster at the meeting.