TORONTO - Radiomics has come a long way from theory to clinical aid for MRI specialists, but standardization and better-quality studies are needed before it becomes a true staple, according to a June 3 presentation at the International Society for Magnetic Resonance in Medicine (ISMRM) annual meeting.
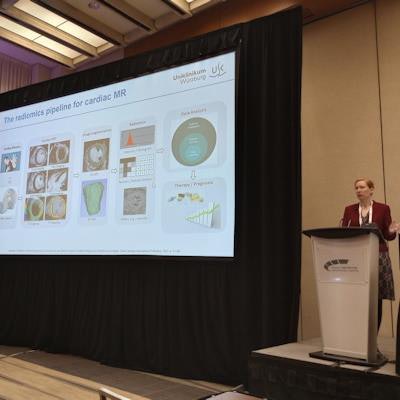
That was the takeaway message for the keynote presentation by Bettina Baessler, MD, from University Hospital Würzburg in Germany.
"There are huge opportunities, and that is why so much research has been put into the field," Baessler told AuntMinnie.com. "It has huge potential, and this has been demonstrated by good-quality studies."
Baessler's research focuses on MRI, radiomics, AI, and machine learning. This includes translating radiomics into clinical practice, validating quantitative imaging techniques, and applying machine learning algorithms in medical imaging.
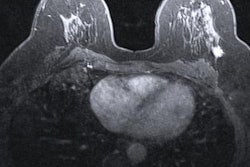
Radiomics has gained in popularity in recent years, with over 7,700 studies published within the past decade. This method takes big data from image and clinical features and compiles them to aid in diagnosis of suspicious findings. The method can be applied to various parts of the body, including for cardiac, lung, and breast imaging among other areas.
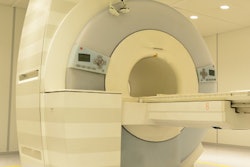
 Bettina Baessler, MD, from University Hospital Würzburg in Germany gives a keynote presentation on the state of MR radiomics at the ISMRM annual meeting in Toronto. Image by Amerigo Allegretto.
Bettina Baessler, MD, from University Hospital Würzburg in Germany gives a keynote presentation on the state of MR radiomics at the ISMRM annual meeting in Toronto. Image by Amerigo Allegretto.However, not all studies in this area add to the cause. Baessler said what she and colleagues have also seen in recent years are poor-quality studies that do not have clinical significance.
"I would be much happier with an exponential rise of good-quality radiomics studies that address real clinical need, and not doing radiomics [just] because we can do it," she said.
Baessler previously led studies examining the effectiveness of radiomics. One study published in 2020 found that CT-based radiomics could accurately detect benign and malignant histopathology on testicular germ cell tumors. Another study published in 2018 found that using texture analysis of T2 MRI maps leads to high sensitivity and specificity for diagnosing acute infarct-like myocarditis.
Using radiomics to deliver better diagnosis of suspicious imaging findings may seem like a daunting task, but it can definitely be done, according to Baessler. Steps in the radiomics process typically include acquiring and processing images, utilizing open-source applications, choosing the parameters that make the most sense, and extracting features.
"Radiomics should be open source, in my opinion," she said.
However, issues such as operator bias and overfitting can be pitfalls for using radiomics. So, how can researchers avoid these when applying radiomics to clinical research? Baeßler said the following should be considered:
- Using the simplest model possible
- Not using too many features, which may skew results
- Using automated approaches to avoid operator bias as much as possible
- Use the most representative correlation clusters
Baessler said for radiomics to take the next big step in radiology, standardization is needed as the reliability of radiomics studies varies based on multiple factors. These include which measures and features are used in respective studies.
"We have to find a means to standardize the image acquisition side, especially for MRI," she told AuntMinnie.com. "For CT, we see similar technical burdens interestingly because CT is more standardized, but even there, we see the same problems."
For now, though, researchers can take steps to include the best-quality data in their studies. Baessler listed some such methods, including implementing better patient selection methods. The Quality Assessment of Diagnostic Accuracy Studies (QUADAS) tool and the CheckList for EvaluAtion of Radiomics (CLEAR) research reporting guideline should also be used.
Baessler is also co-leading the development of a tool called AutoRadiomics, which she and colleagues wrote in a 2022 study can be used as "a comprehensive solution, one of the analysis steps, or an exploratory tool." The study found that AutoRadiomics could successfully create models using radiomics features, with the researchers also reporting no significant overfitting between training and test sets. The framework and code for analysis are publicly available on GitHub.