CHICAGO - Researchers from Johns Hopkins University have found a way to differentiate benign from malignant kidney lesions with great accuracy, solving a problem that has defied solutions since the advent of CT.
Their weapon is texture analysis, an automated way of looking for telltale heterogeneity in lesions. In the right circumstances, the technique can spell the difference between needing to operate and leaving an indolent tumor alone. Can CT texture analysis (CTTA) serve as a quantitative imaging aid when lesions that may or may not be malignant are detected at CT?
Dr. Siva Raman and colleagues from Johns Hopkins' department of radiology applied the investigational technique in a limited fashion, seeking to discriminate several types of similar-appearing renal masses to distinguish between tumor types. They found that they could distinguish between four tumor types reliably.
"In this study, we looked at a group of lesions that are a major dilemma for all radiologists [and that] today are inevitably resected if you can't tell them apart," Raman said in his presentation. "We chose to apply texture analysis to an area where radiologists have struggled since the advent of CT -- and that is our long-term inability to distinguish the most common histologic subtypes of renal masses. ... I think we were able to do better than radiologists could."
Look-alikes on CT
The toughest candidates are clear cell renal cell carcinoma (RCC) and papillary RCC, which are all but impossible to distinguish from benign lesion types such as oncocytomas. Overlap in enhancement characteristics and morphology for individual lesions makes prospective differentiation difficult, Raman said.
Currently, all solid renal masses are surgically resected. But the ability to distinguish lesion types prospectively would have a major impact on treatment, he said.
 Dr. Siva Raman from Johns Hopkins University.
Dr. Siva Raman from Johns Hopkins University.In their study, Raman, Yifei Chen, James Schroeder, PhD, and colleagues used CTTA software from U.K.-based TexRad to retrospectively analyze 20 clear cell RCCs, 20 papillary RCCs, 20 oncocytomas, and 20 Bosniak I renal cysts.
These served as a training cohort, and the group recruited an additional test cohort of 19 patients presenting with five papillary renal cell carcinomas. So in total, there were 80 lesions to train the model and 19 to test it, Raman explained.
All patients underwent preoperative imaging with a three-phase renal mass CT protocol. Because of some limitations in the software, all analyzed lesions had to measure at least 2 cm in all three dimensions.
The researchers manually drew regions of interest (ROIs) around each mass on multiple slices, and each lesion was analyzed using arterial-, venous-, and delayed-phase images.
"We did have noncontrast CT, but we couldn't reliably draw margins around the lesions," Raman said. "And this software, unfortunately, doesn't allow for automated segmentation ... so we actually had to draw ROIs in multiple slices in order to get a range of representative textures."
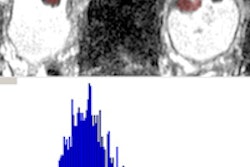
They acquired multiple histographic parameters from the software, including mean gray-level pixel intensity (average brightness), entropy (irregularity), standard deviation of pixel intensity (width of the distribution of intensities), kurtosis (peakedness of distribution), and skewness (asymmetry of distribution).
"Every time we analyzed an ROI, we produced about 32 variables to go into our model," Raman said. "We created multiple classification regression trees ... and each 'tree' was randomly split on a small subset of our input variables."
Unfiltered and filtered images each produced five texture variables, for a total of 30 variables per slice. One predictive model used all three phases, and another used only the arterial phase; altogether, almost two-thirds of the data collected are actually utilized in the predictive model built for the study.
High accuracy for the toughest lesion types
What did they find? "The accuracy rates are very good," Raman said. "The model can essentially stratify all papillary RCCs and all simple cysts, and the error rates for oncocytomas and clear cell RCCs are 11% or less. Now, to give you some perspective on that, there is no radiologist who can distinguish an oncocytoma from a clear cell RCC with any accuracy. I certainly can't, and I do a lot of body imaging."
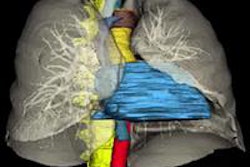
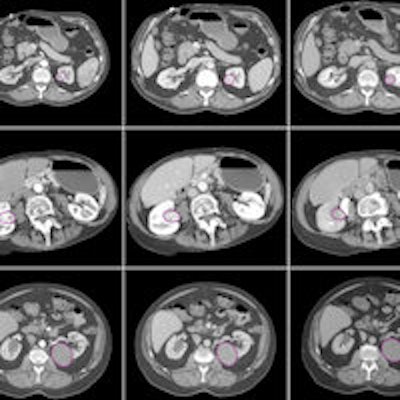
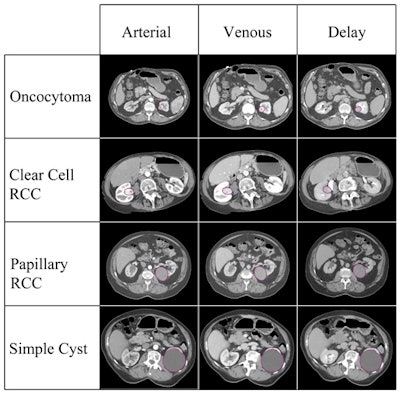 The various lesion types included in the study, imaged in three CT phases. Image courtesy of Dr. Siva Raman.
The various lesion types included in the study, imaged in three CT phases. Image courtesy of Dr. Siva Raman.If the classification of each case were allowed to be the principal class used for all of the slices analyzed, the model that incorporated the three imaging phases together would have correctly classified all 19 lesions. The model using arterial-phase data alone would misclassify a single oncocytoma as a clear cell RCC, correctly categorizing 18 of 19 cases. Finally, the model that used venous-phase data alone would correctly categorize 13 of 19 cases.
Thus the random forest model (the input variables represent the trees) was used to build a predictive model to classify lesion types, Raman said. It was able to correctly categorize clear cell RCCs in 92.5% of cases, papillary RCCs in 100% of cases, oncocytomas in 89% of cases, and cysts in 100% of cases.
"It goes without saying that if you use a model built on a single contrast phase, your accuracies aren't going to be as good, and that's particularly true of models that utilize the venous and delayed phases alone," Raman said. Body imagers know that the arterial phase is the one that provides the best information overall.
"Texture analysis provides quantitative measures of heterogeneity within a region of interest," he said, helping to solve an ongoing dilemma for radiologists.
"Specifically, if you look at our differentiation of oncocytomas and clear cell RCCs, I think we were able to do better with the predictive model than radiologists could" without it, Raman said.
After the talk, a moderator asked whether one could really take the risk of leaving a kidney lesion alone -- no matter how good the model.
"Regardless of how good this model is, I don't think any surgeon is going to leave a 3-cm mass," Raman said. "But I think where you can start using this is in smaller lesions with viable data, and a lot of these lesions are being followed over time. There are data that [indicate] the smaller the lesion, the likelier a mass is benign, in general. So for the smaller lesion, I think you can have a certain level of confidence in following it over time."