Yale University researchers have developed an artificial intelligence (AI) algorithm for classifying liver lesions on MRI scans that explains the reason for its decisions. The algorithm could address concerns about the "black box" nature of AI, according to research presented at last week's RSNA 2018 meeting.
After training a convolutional neural network (CNN) to classify liver lesions, a multidisciplinary group from Yale University in New Haven, CT, found that the deep-learning algorithm yielded higher sensitivity and specificity than the average performance of two experienced radiologists. Importantly, the system can also "explain" its findings to radiologists.
"Our feasibility study offers a path toward more reliable results and may provide diagnostic radiologists and interdisciplinary tumor board members with rapid decision support with verifiable results," co-author Dr. Julius Chapiro told AuntMinnie.com. "Ultimately, it may help us understand the intricacies of complex multiparametric imaging data in liver cancer patients and push the boundaries of patient care quality and efficiency."
A challenging task
Diagnosing liver tumors remains a challenging task and frequently results in inconsistent reporting. The Yale research team, which includes diagnostic and interventional radiologists, biomedical engineers, and oncologists, focuses on developing novel imaging biomarkers and improved paradigms for the diagnosis, treatment, and management of liver cancer, Chapiro said.
AI techniques -- and specifically deep-learning algorithms -- have the potential to substantially improve diagnostic accuracy and workflow efficiency, he said. The intrinsic "black box" nature of deep-learning systems is a major barrier to clinical use, however.
"Without being able to justify and explain the decision-making process of these algorithms, conventional deep-learning systems could act unpredictably and, thus, fail to translate into meaningful improvement of clinical practice," Chapiro said. "Simply put, no one can trust the machine if there is no way to track how it arrived at a specific diagnostic conclusion."
As a result, the team developed a deep-learning algorithm that could provide improved accuracy for diagnosing liver cancer, while also justifying and explaining its decisions to radiologists, he said.
Using 494 different focal hepatic liver lesions in 296 patients, the group trained and tested a CNN to autonomously recognize specific imaging features on raw multiphase contrast-enhanced MRI scans. After recognizing these features, the system then employs a decision-verification algorithm to justify how it arrived at a specific diagnostic decision, such as characterizing a lesion as a hepatocellular carcinoma (HCC), said co-author Clinton Wang. The group has submitted a patent application for their model.
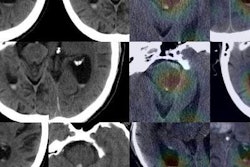
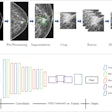
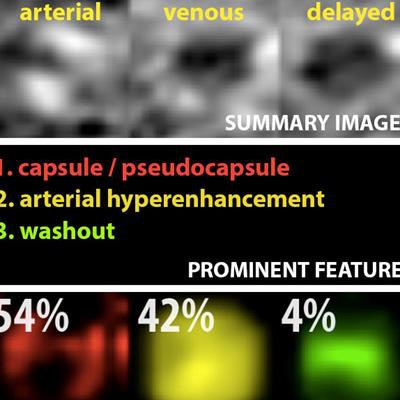
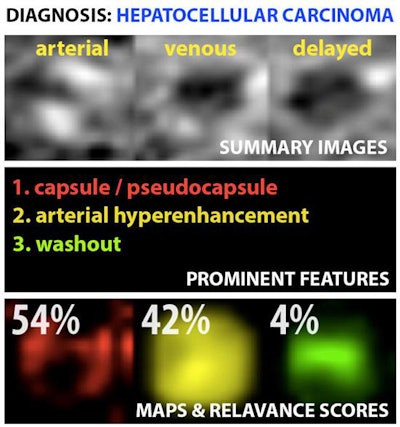 Example output of the Yale group's patent-pending "explainable" deep-learning system. In addition to providing a prediction (e.g., hepatocellular carcinoma), an explainability algorithm justifies the system's decisions with radiological features, feature maps, and relevance scores that contributed to its analysis. Image courtesy of Dr. Julius Chapiro.
Example output of the Yale group's patent-pending "explainable" deep-learning system. In addition to providing a prediction (e.g., hepatocellular carcinoma), an explainability algorithm justifies the system's decisions with radiological features, feature maps, and relevance scores that contributed to its analysis. Image courtesy of Dr. Julius Chapiro.The 494 liver lesions included six liver lesion types: simple cysts, hemangioma, focal nodular hyperplasia, HCC, intrahepatic cholangiocarcinoma, and colorectal carcinoma metastasis. These lesions were defined on T1-weighted MRI scans (arterial, portal venous, and delayed phase) and were divided into a training set of 434 lesions and a test set of 60 lesions. Next, the group compared the algorithm's performance on the test set with that of two board-certified radiologists.
| Diagnostic performance across 6 liver lesion types | ||
| Average of 2 radiologists | Deep-learning algorithm | |
| Sensitivity | 82.5% | 90% |
| Specificity | 96.5% | 98% |
The deep-learning algorithm achieved an overall accuracy of 92%, compared with 80% to 85% for the experienced board-certified radiologists, according to Wang.
"While other published studies have focused on classifying individual 2D slices of CT images, our system takes into account the entire 3D volume of a lesion on the reference-standard contrast-enhanced MRI and is, thus, able to take more features into consideration, which probably translates into higher accuracy," Wang said.
Notably, the system also achieved 90% sensitivity for classifying HCC, compared with an average sensitivity of 65% for the two radiologists.
A decision-support tool
With a runtime of 5 msec per lesion and an accuracy of more than 90% across multiple lesion types, the system could function as a quick and reliable decision-support tool in radiology practice, said Charlie Hamm, who presented the research at RSNA 2018. What's more, it may also improve workflows during interdisciplinary tumor boards, at which diagnostic radiologists are frequently challenged with large amounts of new imaging data and have to make rapid and accurate decisions with immediate therapeutic consequences, he said.
"For simpler cases, implementation could drastically improve time-efficiency by automating preliminary reporting, with an interactive mechanism for radiologists to quickly check whether diagnosis and justification are accurate," Hamm said. "For more complex cases, this could act as a 'second opinion,' helping radiologists improve their accuracy for more challenging lesions."
The researchers envision that the system would be implemented clinically in an approach that synergistically combines the experience and intuition of physicians with the computational power and efficiency of AI systems, said senior author Dr. Brian Letzen. They believe the key missing link for practical clinical translation of deep-learning systems is a way for the systems to explain and justify their decisions, so the researchers included an explainable tool for radiologists, according to Letzen.
"This algorithm interrogates the inner circuitry of the deep-learning system, providing radiologists with specific rationale for each decision," he said.
Letzen noted that while this initial study included more lesions with a classical appearance, the group has since also validated the deep-learning model by training it with more atypical pathologically proven lesions. In addition, the team is concentrating on building larger datasets to continuously enhance the model's accuracy, with the potential of expanding the tool for use with other organ systems outside of the liver, Letzen said. Technical refinements to the deep-learning classification and explainability systems also continue to be made.
"From a translational perspective, we are now exploring options to get this tool into the hands of radiologists for practical incorporation into day-to-day clinical practice," Letzen said.
The research received funding support from the RSNA Research and Education Foundation.