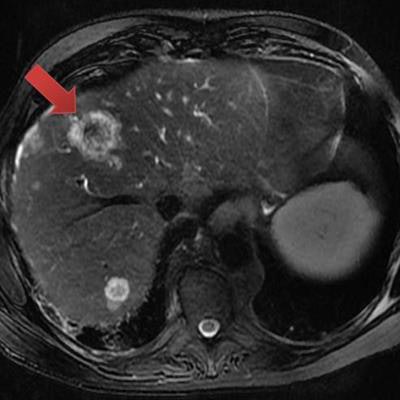
CHICAGO - Machine learning and quantitative MRI features can assist in predicting the KRAS mutation status of tumors in patients with metastatic colon cancer, according to research presented on Wednesday at the RSNA 2017 meeting.
Researchers from Harvard Medical School in Boston found that a machine-learning model that considers tumor radiomic features on MRI could yield a 95% accuracy rate for predicting KRAS mutation status -- a vital factor for determining patient prognosis and treatment decisions in metastatic colon cancer.
"Tumor MRI radiomic analysis may hopefully aid in noninvasively assessing tumor genetic status and informing treatment choices and personalizing therapeutic decisions in patients with colon cancer," said presenter Dr. Dania Daye, PhD.
Guiding treatment decisions
Colorectal cancer is the second leading cause of cancer-related death in the U.S. among men and the third leading cause among women, Daye said. As the treatment options for colon cancer have multiplied, it's becoming increasingly important to improve the decision-making process for who should be treated and how to maximize the treatment risk-benefit ratio, she said.
While many patients with colon cancer are treated with anti-epidermal growth factor receptor (EGFR) therapy, those who have tumors with the KRAS mutation do not typically respond well to this therapy.
"Accordingly, assessment of KRAS mutation status is really essential for prognosis assessment and for guiding treatment decisions," she said.
The Harvard researchers sought to assess the association between quantitative tumor MRI features and KRAS mutation status in patients with metastatic colon cancer. They also wanted to evaluate the role of machine learning-based radiomics in predicting tumor KRAS mutation status, Daye said.
In a retrospective study, the researchers identified 52 patients with stage IV colon cancer who had received contrast-enhanced liver MRI studies at Massachusetts General Hospital (MGH) between 2007 and 2013 for metastatic colorectal cancer to the liver. All patients included in the study had a liver MRI exam at MGH within six months prior to initiating treatment, and they were treated by an oncologist at the MGH Cancer Center. Tumor KRAS mutation status and standard clinical and pathologic prognostic variables were extracted from the medical record.
The patient population had an average age of 57.2; 54% were male and 46% were female. The average lesion size was 3.35 ± 1.9 cm, and the KRAS mutation status was positive in 60% of the patients and negative in 40%.
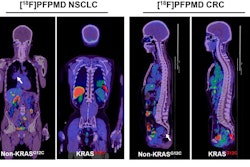
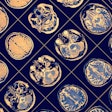
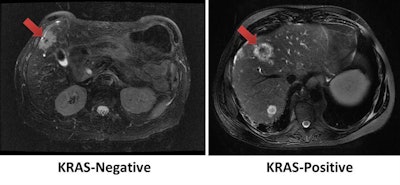 Using tumor radiomic features on MRI, a machine-learning model correctly identified the KRAS mutation status of a KRAS-negative tumor (left) and KRAS-positive tumor (right). Images courtesy of Dr. Dania Daye, PhD.
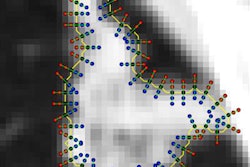
Using tumor radiomic features on MRI, a machine-learning model correctly identified the KRAS mutation status of a KRAS-negative tumor (left) and KRAS-positive tumor (right). Images courtesy of Dr. Dania Daye, PhD.For each patient, the largest hepatic lesion was identified on the portal venous phase T1-weighted fat-suppressed contrast images and manually segmented. The multiparametric radiomic vector was then extracted from each lesion. A total of 52 radiomic features were extracted from each lesion: 20 tumor morphologic features, 15 Laws texture features, 14 Haralick's texture features, and three Tamura texture features.
Univariate logistic regression analysis revealed that six radiomic tumor features were significantly associated with KRAS mutation status. These were circularity, solidity, eccentricity, coarseness, shade, and Gray-Level Co-Occurrence Matrix (GLCM) standard deviation.
Machine-learning model
Next, the researchers used a linear support vector machine (SVM) learning model on all 52 extracted radiomic features from each lesion. The SVM classifier was trained and tested using 10-fold cross-validation to reduce the likelihood of overfitting, Daye said.
The machine-learning model could predict KRAS mutation status with a high level of accuracy, the researchers found:
- Area under the receiver operating characteristic (ROC) curve: 0.95
- Accuracy: 95%
- Sensitivity: 96%
- Specificity: 87%
- Precision: 90%
"Quantitative tumor MRI features exhibit significant association with KRAS mutation status and can be used to predict KRAS status in colon cancer," Daye said.