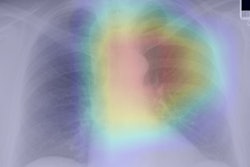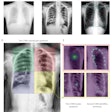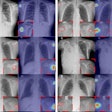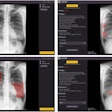
A deep-learning convolutional neural network (CNN) algorithm can estimate skeletal maturity on pediatric hand radiographs as accurately as expert radiologists -- and nearly as accurately as a commercial computer-aided diagnosis (CADx) software application, according to research published online in Radiology.
After training a CNN using more than 14,000 clinical radiographs, researchers led by Dr. David Larson of Stanford University found that the network more than held its own in comparison with three expert radiologists and the bone age assessment provided on the clinical report. The algorithm also only slightly lagged behind the published performance of commercial diagnosis software in the literature.
"Bone age is one of the first applications [that] can be fully automated [using] a model that could be replicated from scratch in a matter of a few days of effort, based on the number of images already present in the PACS of most large radiology practices," Larson said.
An ideal first project
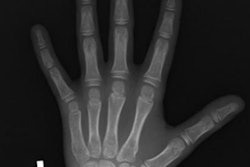
Bone age estimation is a critical task for determining developmental status and predicting ultimate height in pediatric patients, particularly those with growth disorders and endocrine abnormalities. Radiologists can manually estimate patient age on a radiograph, but this process is considered to be tedious, time-consuming, and prone to inter- and intrareader variability.
 Dr. David Larson of Stanford University School of Medicine.
Dr. David Larson of Stanford University School of Medicine.Larson's deep-learning initiative began more than a year and a half ago when co-author Matt Chen, then a master's student, was seeking a real-world project for his computer vision class. Bone age assessment seemed to be an ideal first project, as it involves a single clinical question based on a single image type, with a quantitative output that can be directly compared with human performance, Larson said.
After Chen achieved good results using the publicly available digital hand atlas developed by the University of Southern California, the researchers wanted to see if it was possible -- if they could get enough images -- to develop a bone age assessment model that could top an expert human reviewer, according to Larson.
Using a dataset of more than 14,000 clinical radiographs of the left hand from Lucile Packard Children's Hospital at Stanford and Children's Hospital Colorado in Aurora, CO, the researchers trained and tested four different deep CNN models. These models were trained using different numbers of images (1,558, 3,141, 6,295, and 12,611 images, respectively) to evaluate the effect of the size of the training set on the algorithms' performance. Of the initial dataset, 90% of the images were used for training and 9% were reserved for validation of the models.
Before being used to train the algorithms, the images were converted to DICOM format, downsized to a resolution of 256 x 256 pixels, and enhanced with contrast-limited adaptive histogram equalization. They were then further cropped to a resolution of 224 x 224 to accommodate transformations on the augmented images such as random flips, crops, and contrast adjustments, according to the researchers (Radiology, November 2, 2017).
Strong performance
After training was complete, the researchers compared the best-performing model's performance on 200 radiographs from the test set with the performance of four expert human reviewers -- three study co-authors and the actual clinical report.
"We found that the model performed better than the clinical report and one of the reviewers and not significantly different than the other two reviewers," Larson said.
The authors found a mean difference of 0 years between the model's bone age estimates and the reviewers' estimates. The mean root mean square (RMS) -- another measure of performance -- was 0.63 years and the mean absolute difference was 0.5 years.
"The estimates of the model, the clinical report, and the three reviewers were within the 95% limits of agreement," they wrote.
Not surprisingly, the researchers found that the most accurate model was the one trained with the most images:
- Training set with 1,558 images -- RMS age difference: 1.08 years
- Training set with 3,141 images -- RMS age difference: 0.91 years
- Training set with 6,295 images -- RMS age difference: 0.78 years
- Training set with 12,611 images -- RMS age difference: 0.73 years
The improvement in accuracy started to level off fairly quickly, however, Larson said.
"For example, a model that was trained with only half the number of images saw only a 6% decrement in performance," he told AuntMinnie.com. "This suggests that we might be able to use relatively small dataset sizes to train deep-learning models."
Comparison with BoneXpert
In a separate test on 1,377 examinations from the digital hand atlas, the researchers found that their model had an RMS age difference of 0.73 years, compared with 0.61 years -- as published in the literature -- for the commercially available BoneXpert software (Visiana). Larson pointed out that this result shows that deep learning isn't necessarily better -- especially for very focused and reproducible tasks -- than the feature extraction techniques used traditionally in CADx software.
"The difference is that once we got a hold of the dataset and made a few tweaks, our model took a few weeks to develop using open-source software, compared to years of painstaking work by the developers of BoneXpert," he said. "This illustrates an important point: Deep learning not only has the potential to create powerful diagnostic tools for radiology, [but also] these tools can be created by virtually anyone with a small investment of time and a relatively low number of training images."
Current update
The researchers are now developing an application to fully automate the model and integrate it into the workflow -- from image acquisition to integration with the clinical report, Larson said.
In addition, they donated the training dataset to the RSNA to serve as the basis for the recent RSNA Pediatric Bone Age Challenge. Nearly 300 entrants competed in the contest to develop an algorithm that could most accurately determine skeletal age on the pediatric hand radiographs.
The winners will be announced at RSNA 2017 in Chicago, "but the organizers have already announced that the winners' models were better than our model, which is exactly what we were hoping," Larson said.