Top Story
Latest News
Cases of the Week
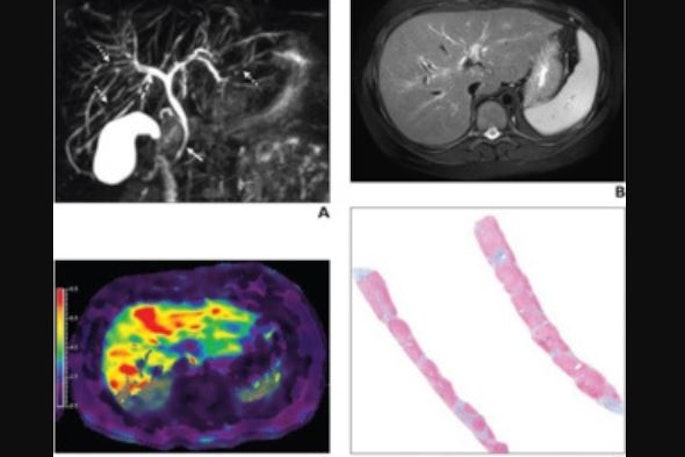
Check out our Cases of the Week!
More from AuntMinnie
Evergreen Theragnostics raises $26M
April 19, 2024
Segmed partners with Beacon Health System in Indiana
April 19, 2024
Accuray opens training facility in Switzerland
April 18, 2024
Lumicell's Lumisight, LumiSytem get FDA nods
April 18, 2024
Radiology Leadership Institute names award recipients
April 18, 2024
GE HealthCare launches two new ultrasound systems
April 18, 2024
AdvaMed urges Congress prioritize R&D tax credit
April 17, 2024
Swoop system tested in Alzheimer’s patients
April 17, 2024
PET tracer for gliomas under expedited review
April 17, 2024